Supported Applications
FastQC
-
Description
a quality control tool for high throughput sequence data.
-
Usage
To list all executables provided by FastQC, run:$ biogrids-list fastqc -
Installation
Use the following command to install this title with the CLI client:$ biogrids-cli install fastqcAvailable operating systems: Linux 64, OS X INTEL -
Citation Note
Users of the program should cite the url http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc -
Webinars
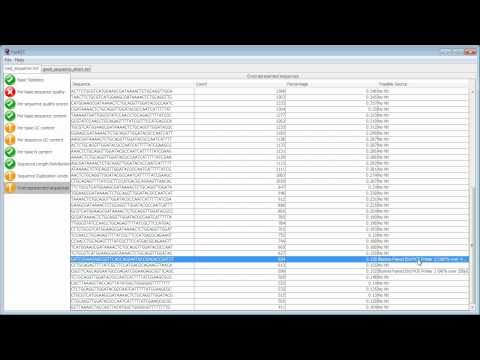
FastQC is an application which reads raw sequence data from high throughput sequencers and runs a set of quality checks to produce a report which allows you to quickly assess the overall quality of your run, and to spot any potential problems or biases.
This video demonstrates the use of FastQC to analyse some sequence data and goes through the results to explain what a good dataset looks like, and what sort of problems you might encounter.
-
Keywords
ChIP-Sequencing, DNA-Sequencing, High-throughput sequencing, Read Quality Control, RNA-Sequencing, WGS Analysis
-
Default Versions
Linux 64: 0.11.9 (10.8 MB)
OS X INTEL: 0.11.9 (124.1 MB) -
Other Versions
Linux 64:
0.11.2 (1.7 MB) , 0.11.5 (10.6 MB) -
OS X INTEL:
0.11.5 (10.8 MB)
Developers
Simon Andrews