SBGrid Consortium homepage

With Support from the HMS Tools and Technologies Committee and NSF EAGER#1448069. BioGrids now available at Harvard Medical School and Boston Children's Hospital. This project is a part of the MOC alliance.

Read our September newsletter
Latest upgrades from BioGrids, including latest software updates, new members and special events. Published montly by BioGrids.

Capsules Technology
We have recently published a manuscript describing our Capsules technology used to deploy SBGrid and BioGrids software collections. Read more in Acta Cryst Section D, special 75th ICUR anniversary issue, …
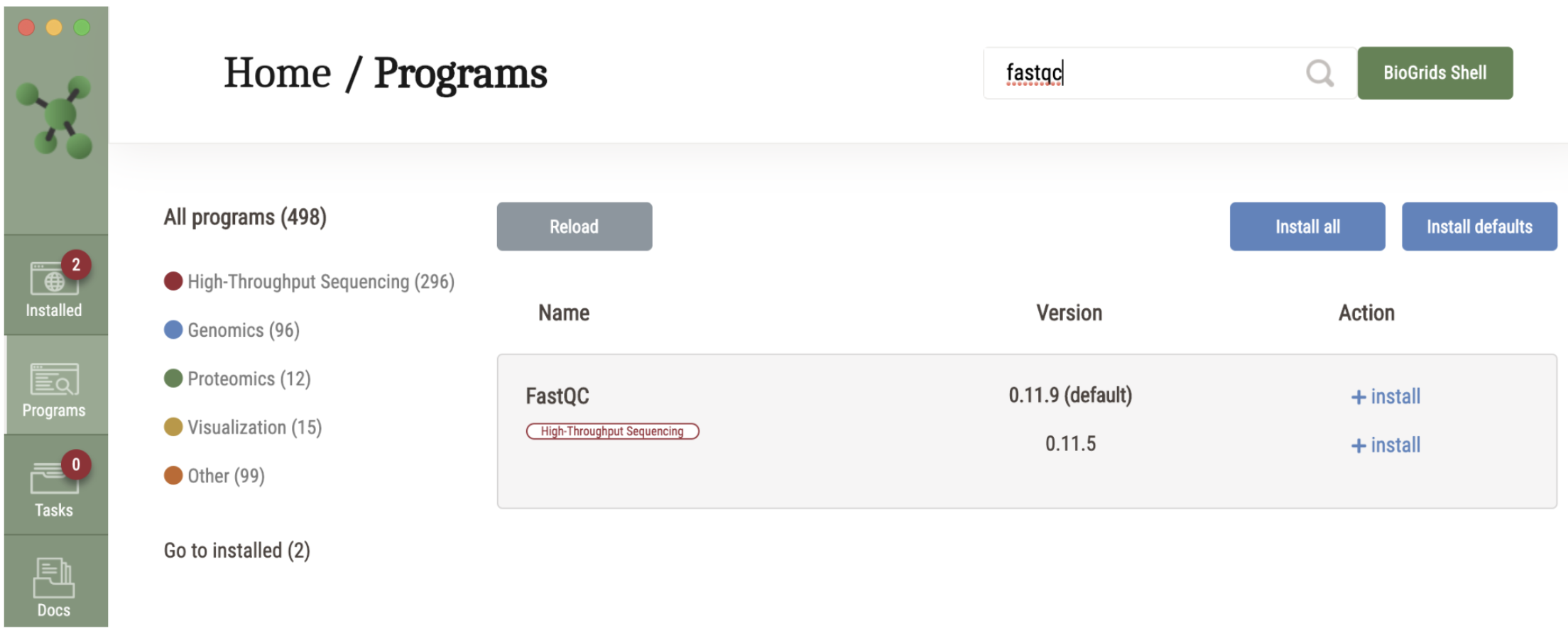
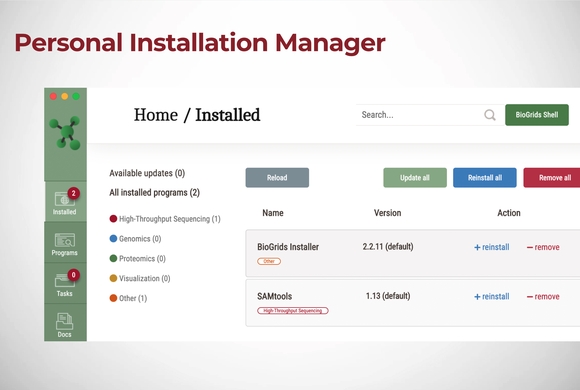
BioGrids for SBGrid Users Tutorial
For SBGrid users who are already familiar with the SBGrid software installation platform, members may opt-in in to include the BioGrids software stack in their installation. BioGrids in an SBGrid-style …

Training Opportunities at HMS
A range of bioinformatics training opportunities are available to HMS researchers, with classes ranging from 2-hour introductory Python lectures to a 10-day in-depth NSG data analysis courses. Check out the …
-
Report Software Bug
Software fixes, depending on priority take 1-14 business days to complete.
-
Request New Software
New software packages are added on a monthly basis, or more often when requested.
-
Other Support
Any other requests or comments can be submitted here.
-
Grant Support
A statement for your resources section, support letters, and how to cite SBGrid.